How to make a cross-functional team thrive and what value it brings
Learn how to build a cross-functional team and how they can spur innovation, foster collaboration, and accelerate problem-solving in your business.

A long, long, long time ago programmers used to write code, compile it, and then analyze its results afterwards. The old-time dream was about more interactivity when working with the software, having the ability to experiment and being able to see the results immediately.
Spreadsheets showed business users how easy it was to try different formulas or macros to automate repetitive tasks, and to sort, group and visualise data. It all happened immediately without waiting for the developers to change the code, compile it, build it, or deploy it. According to Microsoft, Excel macros are the most popular programming languages in the world. And I agree, Office apps are where the first citizen developers were born.
Data science is not as deterministic as traditional software development and data scientists, as people, come from different backgrounds other than API and frontend developers. Creating models is an experimentation based on knowledge and experience, tuning models, exercising machine learning techniques, trying different datasets, manual data cleaning, parameters tuning, and seeing the results, all in a constant feedback loop.
That’s why data scientists are in love with their computational notebooks, with Jupyter Notebook being the most popular one.
The UI of the notebook is a web page with cells. The cells are key here and can consist of a description, code, and operation results. Once the call is added, it is executed and the result is immediately visible. You can go back as many steps as you want, even change the code or data, and then see what are the results of the changes.
In the background, there’s a notebook server running, called a kernel, which executes commands and generates outputs. So, notebooks always have the server part, no matter if it’s deployed locally (localhost) or over the network.
Below is an example of Notebook from our Julia language research which you can find on our GitHub repository.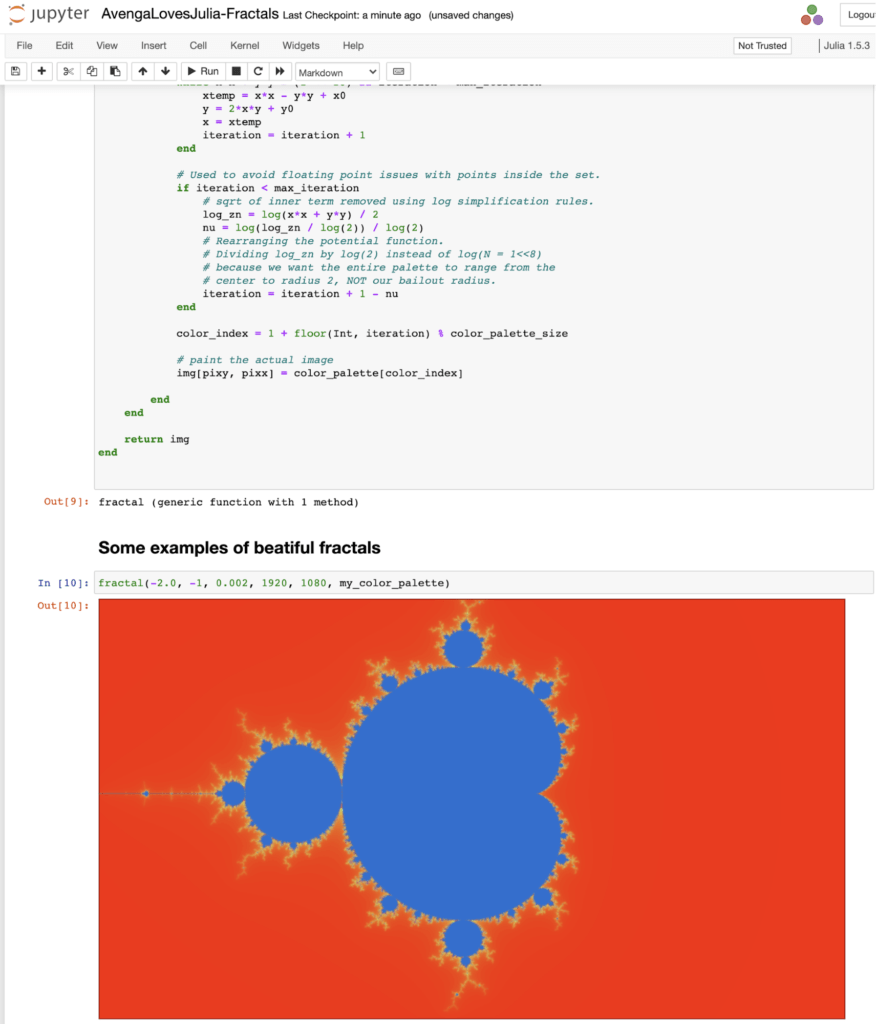
As you can see, notebooks are a very interactive and an immediate way of trying things.
Generating images and graphs on the fly with plot command and other visualization libraries is also a great way to see what data looks like, before and after the transformation.
It builds confidence that the operations and models are correct and will generate good results; also in a full data set.
Manual tweaking is much faster than a full cycle of building, deploying and testing. Ideas are verified quickly, ad hoc. When something doesn’t work as expected it’s often enough to go back just one step to correct things.
Debugging is considered easy, because it only takes a moment to look and determine visually if the resulting graph or table looks ok.
The results of operations are saved in the notebook itself, so it saves a lot of time that would normally be needed to regenerate partial results from the previous steps.
The results of the steps are preserved so when someone downloads it, the notebook is able to see everything (data, code, results, images) without the need to recreate and run all the steps.
Notebooks are easier to understand, self-learn, and try than full fledged applications in Python, R or Julia.
They represent a high level abstraction, so one does not need to worry about Python’s version and configuration, underlying operating system, etc.
It’s self documenting, so the code is a near explanation of what it is doing, then the results follow and flow to the reader; and someone learning can be very productive.
That’s why Julia language research, for instance, was a natural choice for Avenga Labs.
Notebooks are great for demos, showcasing a hypothesis and documenting the validity of the approach.
→Read about Avenga Data science’s perspective on COVID-19: a real life example
Notebooks are a great tool for what they were designed to do, which is an ad hoc experimentation. The primary environment is, naturally, the laptops and workstations of data scientists and analysts, but then running notebooks on dedicated servers. However, recently there are new options to deploy notebooks into Kubernetes clusters.
Data scientists and DevOps teams are even encouraged to do so by solutions for Kubeflow (MLOps pipeline), Databricks and others. The message is that with a few changes, your notebook will run just fine with Kubernetes. And when it runs inside the Kubernetes cluster, it must be modern and good. Some even consider deploying computational notebooks into the Kubernetes cluster as the realization of MLOps / DataOps dreams.
→Explore why DataOps – more than DevOps for data
And it’s probably here, where the message from cloud vendors ends on a positive note, but it is very much one sided. We’re better than that though.
From the programmer’s perspective, computational notebooks represent the typical differences between scripting languages and compiled languages (for instance Python vs. Java). So they are better for trying things, but much worse for the production environments.
There are many who oppose deploying notebooks into the cloud native environments. There are even stronger voices saying that deploying notebooks into production environments of any kind is a very bad idea. This seems to have become a trend based on the popularity of opinions.
Let’s go through some of the well known limitations and disadvantages of computational notebooks.
Notebooks are very easy to try things out in and to visualise the results immediately. It makes them easy to be overused for testing and thus replacing more reliable and industrialized unit tests. What is easy for a single person working with notebooks to do does not scale well and work with continuous delivery and deployment pipelines.
One bad habit is long code in cells that do multiple things at the same time. It does work but it’s not structured properly. This happens all the time because in the experimentation stage the code quality and decomposition seem to not be important.
This kind of code, which is sometimes hundreds of lines of functions, local variables, etc., is very hard to read by other developers and even harder to understand and maintain. The next cells depend on the previous cells and visual feedback becomes a double-edged sword. It makes data scientists less imaginative with more complex, harder to visualise in 2D/3D scenarios.
Versioning of the notebooks is really hard compared to the traditional languages in the Git repository. Yes, the notebook files can be shared, but merging changes, versioning, branching, and all the things developers do with code in large projects is much more difficult, and often infeasible from a pragmatic standpoint.
They are good for sharing the final results of experimentation, to a much lesser extent than working collaboratively on them.
Code autocompletion, linting, and other time saving features are missing.
There’s much less need to look up how the function works in the other window in IDEs than in Notebooks. With notebooks, it’s almost a constant searching for code samples and function descriptions in one tab while writing code in another tab.
Debugging is easy, but very limited, and there are no proper breakpoints.
Interactive convenience is different from production code, which is supposed to be as autonomous as possible, especially in the spirit of everything as code and total automation.
Data safety when sharing, can be jeopardized, as notebooks may contain data samples from actual data sources, resulting in sensitive data being exposed unexpectedly.
Scaling is very hard to do in notebooks. Code modularization is also difficult to achieve. The separation of concerns and other best practices from application development are, again, not easy to implement. It’s almost impossible to talk about the proper architecture, etc.
This limitation is well known in the data industry and new versions of tools promise to address these limitations, but with varying degrees of success and the whole spectrum of opinions about them. For some, especially working in very small teams, there’s almost no problem, but for others the current state of the toolset is unacceptable.
The current strong trend for MLOps (Machine Learning Operations) and DataOps is about bringing the best practices from modern software development techniques into data projects as well.
Throwing computationals notebooks away is neither a good idea nor possible, given their great benefits and the entire work culture around them.
Data scientists love their notebooks, models, and tuning as well as the efficiency of the results of instantaneous feedback. There is also the ability to tinker with models, code and visualisation. Not many data scientists like programming and they are often not the best coders, which is why they have chosen different career paths; and they shine in different areas than application programmers.
Yet, the times are changing, whether they like it or not.
Automated testing, proper version control, automated deployment, and reduced dependency on individual team members enables better scalability and predictability. And, data projects should no longer be excluded either.
The problem with this approach is that it also requires a deep cultural shift from the traditional data science approach to industrialized software development. Data scientists are about to learn the skills of software developers, and learn about the DevOps requirements for modern deployment pipelines.
Right now there seems to be no other way around than the close collaboration between data scientists, developers and DevOps. All sharing their skills with each other to learn about their point of view and figuring out the optimal solution.
There are pros and cons of computational notebooks and the key is to use them where they are the most effective and switch to other tools for the modern delivery pipelines.
It’s part of full cycle development and the developers movement, which enables the faster and more predictable development of any type of solutions as well as data platforms.
The journey is not easy and the benefits won’t be visible tomorrow, but the sooner you start it the better it will be. We, at Avenga, are ready and open to help as you adopt this shift in your context.
* US and Canada, exceptions apply
Ready to innovate your business?
We are! Let’s kick-off our journey to success!